Protein Dynamics
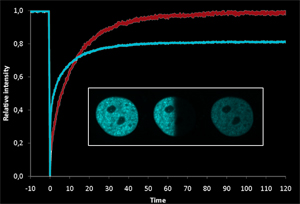 Epigenetic factors function in a complex network with changes in time and location. Using FRAP/FLIP protein kinetics can be studied at a specific time point during the cell cycle or at a specific location in the cell. Furthermore, in these vivo experiments the proteins are analyzed without neglecting chromatin structure, protein modifications or interacting factors.
Epigenetic factors function in a complex network with changes in time and location. Using FRAP/FLIP protein kinetics can be studied at a specific time point during the cell cycle or at a specific location in the cell. Furthermore, in these vivo experiments the proteins are analyzed without neglecting chromatin structure, protein modifications or interacting factors.
These methods allow us to compare the dynamics of the wild type protein with mutants or with different cellular contexts. In order to extract absolute values, we are using kinetic modeling to extract diffusion coefficients or association rates. We are focusing on factors related to DNA methylation like the DNA methyltransferase 1 (Dnmt1), Dnmt3a/b and Uhrf1/2.
A variation of these techniques is Microirradiation. Irradiation with a 405 nm laser induces DNA damage at specific areas in the nucleus. Enrichment of proteins at these sites provides information about the dynamics of DNA damage responses.
Publications:
Cardoso MC, Schneider K, Martin RM and Leonhardt H (2012). Structure, function and dynamics of nuclear subcompartments. Curr Opin Cell Biol., 2012 Jan 5. [Epub ahead of print]
Pichler G, Wolf P, Schmidt CS, Meilinger D, Schneider K, Frauer C, Fellinger K , Rottach A and Leonhardt H. Cooperative DNA and histone binding by Uhrf2 links the two major repressive epigenetic pathways. J. Cell Biochem. Epub 2011 May 19.
Rottach A, Frauer C, Pichler G, Bonapace IM, Spada F and Leonhardt H (2010). The multi-domain protein Np95 connects DNA methylation and histone modification. Nucl. Acids Res. 38, 1796-804.