Technology Development
Antibody generation
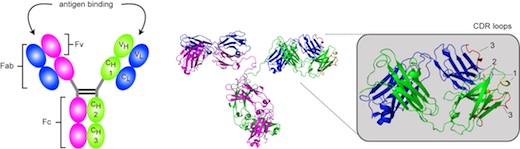
Antibodies are powerful tools to study protein-protein interactions, protein localization and posttranslational protein modifications in their natural cellular environment. Over the past decade, we successfully generated in close collaboration with Dr. Elisabeth Kremmer (Helmholtz Zentrum München) several high quality monoclonal antibodies (rat and mouse) that are suited for cell biological and biochemical applications. Besides polyclonal antibodies, we will include in our portfolio small nanobodies that are generated in camels and are able to detect their specific antigen in living cells and in real time. Using these antibodies in high throughput imaging and biochemical assays, we will be able gain more insights in the epigenetic protein network.
Designer TAL-Effectors
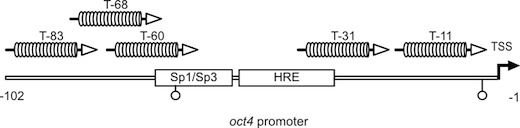
The ability to specifically manipulate the expression of endogenous genes by engineered designer transcription factors has wide-ranging applications in basic and applied biology. Availability of suitable DNA binding scaffolds that can be tailored to bind user-defined target sequences has been the major limitation in the generation and application of designer transcription factors. Recent studies however demonstrated that transcription activator-like effector proteins (TALEs) from the plant pathogenic bacterial genus Xanthomonas contain a DNA binding domain that can be adjusted to bind any desired target sequence with high specificity. The TALE DNA binding domain is composed of tandem arranged 33-35 amino acid repeats, with each repeat binding to one base. Base preferences of repeats are specified by residues 12 and 13, known as the repeat variable diresidues (RVDs), that determine preferential pairing with A (NI), C (HD), G (NK) and T (NG) nucleotides, respectively. The use of this TALE code facilitates the assembly of TALE repeat arrays that bind any desired DNA sequence. In collaboration with the group of Prof. Dr. Thomas Lahaye we successfully developed a method for targeted transcriptional activation using designer TALEs. We are currentlyworking on new methods to further exploite the full potential of this new technology.

